Note
Go to the end to download the full example code
Generating 1D scatter plots with outliers¶
This examle will plot the raw data points in a 1D scatter plot. Here, we attempt to highlight outliers by calculating the interquartile range.
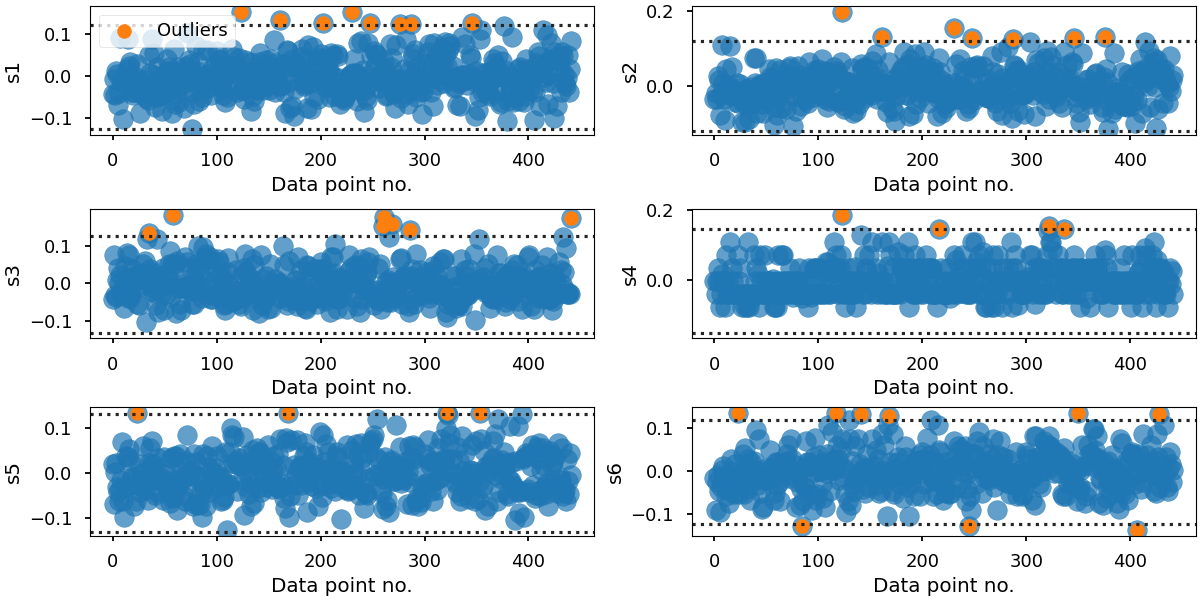
Possible outliers for "s1":
| | age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 |
|-----|---------|--------|---------|---------|------|-------|----------|---------|-------|---------|
| 123 | 0.0054 | 0.051 | 0.035 | -0.0011 | 0.15 | 0.2 | -0.062 | 0.19 | 0.016 | 0.073 |
| 161 | -0.045 | 0.051 | 0.064 | 0.07 | 0.13 | 0.13 | -0.04 | 0.11 | 0.076 | 0.086 |
| 202 | 0.082 | 0.051 | 0.0013 | 0.036 | 0.13 | 0.091 | 0.019 | 0.034 | 0.084 | -0.03 |
| 230 | -0.038 | 0.051 | 0.071 | -0.057 | 0.15 | 0.16 | 0.00078 | 0.072 | 0.05 | 0.069 |
| 248 | -0.042 | -0.045 | 0.048 | 0.06 | 0.13 | 0.13 | -0.025 | 0.11 | 0.064 | 0.04 |
| 276 | 0.013 | -0.045 | 0.026 | 0.063 | 0.13 | 0.092 | 0.063 | -0.0026 | 0.058 | -0.022 |
| 287 | 0.045 | -0.045 | -0.0062 | -0.016 | 0.13 | 0.13 | 0.019 | 0.034 | 0.032 | -0.0052 |
| 346 | 0.0054 | 0.051 | 0.018 | 0.032 | 0.13 | 0.13 | -0.021 | 0.071 | 0.063 | 0.015 |
Possible outliers for "s2":
| | age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 |
|-----|---------|--------|---------|---------|------|------|----------|-------|-------|---------|
| 123 | 0.0054 | 0.051 | 0.035 | -0.0011 | 0.15 | 0.2 | -0.062 | 0.19 | 0.016 | 0.073 |
| 161 | -0.045 | 0.051 | 0.064 | 0.07 | 0.13 | 0.13 | -0.04 | 0.11 | 0.076 | 0.086 |
| 230 | -0.038 | 0.051 | 0.071 | -0.057 | 0.15 | 0.16 | 0.00078 | 0.072 | 0.05 | 0.069 |
| 248 | -0.042 | -0.045 | 0.048 | 0.06 | 0.13 | 0.13 | -0.025 | 0.11 | 0.064 | 0.04 |
| 287 | 0.045 | -0.045 | -0.0062 | -0.016 | 0.13 | 0.13 | 0.019 | 0.034 | 0.032 | -0.0052 |
| 346 | 0.0054 | 0.051 | 0.018 | 0.032 | 0.13 | 0.13 | -0.021 | 0.071 | 0.063 | 0.015 |
| 376 | -0.0019 | -0.045 | 0.068 | -0.0057 | 0.12 | 0.13 | -0.025 | 0.087 | 0.046 | -0.0011 |
Possible outliers for "s3":
| | age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 |
|-----|--------|--------|---------|--------|---------|---------|------|--------|----------|---------|
| 35 | 0.049 | 0.051 | -0.031 | -0.049 | 0.049 | -0.0041 | 0.13 | -0.054 | 0.021 | 0.02 |
| 58 | 0.042 | -0.045 | -0.064 | 0.036 | 0.012 | -0.058 | 0.18 | -0.076 | -0.00061 | -0.051 |
| 260 | 0.042 | -0.045 | -0.0084 | -0.057 | 0.0081 | -0.031 | 0.15 | -0.076 | -0.08 | -0.018 |
| 261 | 0.049 | -0.045 | -0.042 | 0.1 | 0.036 | -0.026 | 0.18 | -0.076 | -0.013 | 0.015 |
| 269 | 0.009 | -0.045 | -0.032 | -0.026 | 0.042 | -0.01 | 0.16 | -0.076 | -0.012 | -0.038 |
| 286 | -0.038 | -0.045 | -0.055 | -0.078 | -0.033 | -0.086 | 0.14 | -0.076 | -0.019 | -0.0052 |
| 441 | -0.045 | -0.045 | -0.073 | -0.081 | 0.084 | 0.028 | 0.17 | -0.039 | -0.0042 | 0.0031 |
Possible outliers for "s4":
| | age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 |
|-----|---------|--------|-------|---------|-------|--------|--------|------|-------|-------|
| 123 | 0.0054 | 0.051 | 0.035 | -0.0011 | 0.15 | 0.2 | -0.062 | 0.19 | 0.016 | 0.073 |
| 216 | 0.013 | 0.051 | 0.036 | 0.049 | 0.053 | 0.074 | -0.069 | 0.15 | 0.046 | 0.049 |
| 322 | 0.024 | 0.051 | 0.062 | 0.062 | 0.025 | -0.036 | -0.091 | 0.16 | 0.13 | 0.082 |
| 336 | -0.02 | -0.045 | 0.085 | -0.037 | 0.092 | 0.089 | -0.062 | 0.15 | 0.081 | 0.053 |
Possible outliers for "s5":
| | age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 |
|-----|---------|--------|--------|--------|-------|---------|---------|-------|------|--------|
| 23 | 0.045 | 0.051 | 0.061 | 0.031 | 0.029 | -0.047 | -0.054 | 0.071 | 0.13 | 0.14 |
| 169 | -0.0019 | -0.045 | -0.027 | 0.049 | 0.059 | -0.016 | -0.047 | 0.071 | 0.13 | 0.02 |
| 322 | 0.024 | 0.051 | 0.062 | 0.062 | 0.025 | -0.036 | -0.091 | 0.16 | 0.13 | 0.082 |
| 353 | -0.053 | -0.045 | -0.056 | -0.037 | 0.089 | -0.0032 | 0.0081 | 0.034 | 0.13 | 0.0031 |
Possible outliers for "s6":
| | age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 |
|-----|---------|--------|--------|---------|--------|--------|---------|---------|--------|-------|
| 23 | 0.045 | 0.051 | 0.061 | 0.031 | 0.029 | -0.047 | -0.054 | 0.071 | 0.13 | 0.14 |
| 84 | 0.0018 | -0.045 | -0.04 | -0.1 | -0.029 | -0.03 | 0.045 | -0.05 | -0.068 | -0.13 |
| 117 | 0.06 | -0.045 | -0.021 | 0.087 | 0.045 | 0.032 | -0.047 | 0.071 | 0.079 | 0.14 |
| 141 | -0.027 | -0.045 | 0.048 | -0.047 | 0.034 | 0.057 | -0.08 | 0.13 | 0.045 | 0.13 |
| 168 | 0.0018 | 0.051 | 0.06 | -0.0022 | 0.062 | 0.063 | -0.058 | 0.11 | 0.069 | 0.13 |
| 245 | -0.027 | -0.045 | -0.035 | -0.03 | -0.057 | -0.059 | 0.03 | -0.039 | -0.05 | -0.13 |
| 350 | -0.027 | 0.051 | 0.061 | 0.11 | 0.012 | -0.018 | -0.0029 | -0.0026 | 0.07 | 0.14 |
| 406 | -0.056 | -0.045 | -0.081 | -0.085 | -0.037 | -0.037 | 0.034 | -0.039 | -0.056 | -0.14 |
| 428 | 0.049 | 0.051 | 0.089 | 0.087 | 0.036 | 0.022 | -0.025 | 0.034 | 0.066 | 0.13 |
from matplotlib import pyplot as plt
import pandas as pd
from sklearn.datasets import load_diabetes
from tabulate import tabulate
from psynlig import generate_1d_scatter
plt.style.use('seaborn-talk')
data_set = load_diabetes()
data = pd.DataFrame(data_set['data'], columns=data_set['feature_names'])
variables = ['s1', 's2', 's3', 's4', 's5', 's6']
kwargs = {
'scatter': {
'marker': 'o',
's': 200,
'alpha': 0.7,
},
'scatter-outlier': {
's': 100,
'marker': 'o',
'label': 'Outliers',
},
'figure': {'figsize': (12, 6)},
}
_, _, outliers = generate_1d_scatter(
data,
variables,
show_legend=True,
outliers=True,
**kwargs,
)
for var, out in outliers.items():
print('\nPossible outliers for "{}":\n'.format(var))
data_out = data.iloc[out, :]
print(
tabulate(data_out, tablefmt='github', headers='keys', floatfmt='.2g')
)
plt.show()
Total running time of the script: ( 0 minutes 0.810 seconds)